RNASeq is a Client-Desktop Application of the GPRO suite with custom dedication to manage and run pipelines and workflows for differential expression and enrichment analysis based on RNASeq data using a protocol based on the State-of-the -Art (Fig.1). The application is coupled with an infrastructure of server-side dependencies (pipelines, databases and tools) that we distribute in a container that can be installed on a remote server or on a PC with sufficient RAM. The app also includes a File Transfer Protocol system (FTP) to facilitate the upload and download of files from the user’s computer to or from the server; a progress tracker (job tracking system), and two different execution modes (a “step-by-step” mode, and a “pipeline-like” mode).
The Step-by-Step mode is a procedure similar to those implemented in Galaxy (Afgan et al 2018) and others GUI-based solutions for NGS data analysis. This mode organizes the different steps of the RNASeq workflow (i.e. quality analysis, preprocessing, mapping, transcriptome assembly and/or quantification, differential expression, and enrichment) into an intuitive menu providing a selection of command line interface (CLI) third party software for each step allowing different protocols to perform the analysis with or without a reference genome or transcriptome. At the same time each CLI tool has an interface implementation with distinct fields to declare the inputs and outputs files or for tuning the options and parameters provided by that tool.
In contrast, the pipeline mode is a pipeline configuration system allowing the user to execute all the steps of a given protocol automatically one after the other. To this end the user just need to select an specific pipeline from a list, declare the experiment design as well as the input and output data, then configure the option and parameters and finally run the pipeline where the distinct analyses will be executed sequentially one after another.
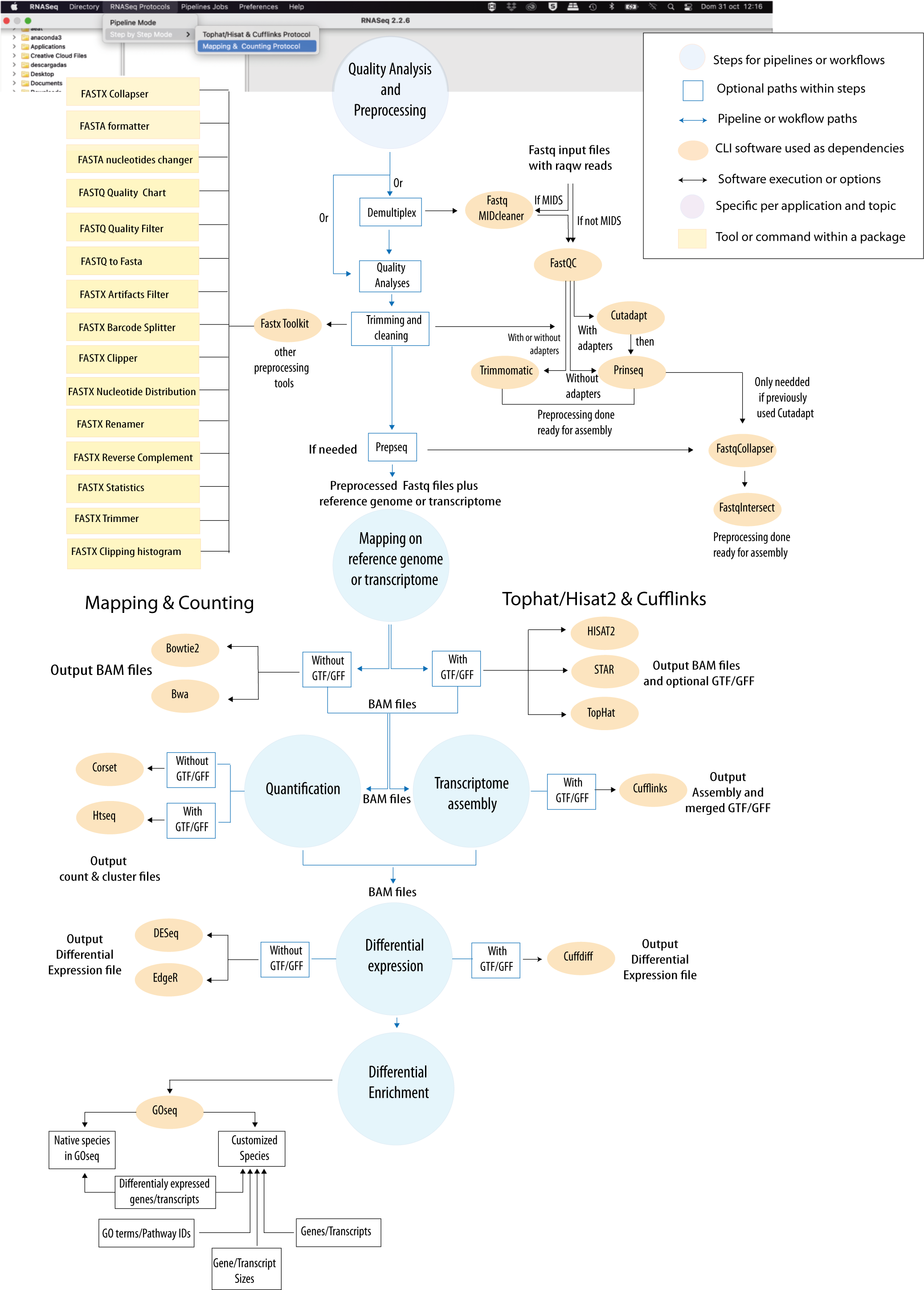
Figure 1: Bioinformatic protocol implemented in RNAseq for differential expression and enrichment analysis according to the most common RNA-seq practices. The tool provides two execution modes (Step-By-Step and Pipeline-like) and two alternative analytical paths (“Mapping & Counting” and “Tophat/Hisat2 & Cufflinks”). An interactive version of this protocol is available at GENIE our virtual assitant.
java -version |
$ java -version
|
RNASeq-win32.win32.x86_64.zip file and unzip it.
Then browse to the executable file “RNASeq.exe” and execute/run it.
RNASeq-macosx.cocoa.x86_64.zip file and unzip it. Then browse to the executable binary file "RNASeq.app" and execute/run it.
RNASeq-linux.gtk.x86_64.zip file and unzip it. Then browse to the executable binary file “RNASeq” and execute/run it.
RNASeq is a Client Side + Server Side solution thus meaning that the application is coupled via API with a bioinformatic infrastructure called GPRO Server Side that contains all the dependencies needed by RNASeq to execute the workflows and pipelines. These dependencies are scripts, databases and the following third party CLI software:
The GPRO Server Side can be installed in the PC of the user or in remote servers as a Cloud Computing resource. However, its installation is a complex task due to the lot of dependencies and requirements (besides of the CLI software) for installing and running this infrastructure. For this reason, we distribute the GPRO Server Side in a Docker container that can be easily installed for the user in a couple of steps. Indications for installation of the GPRO Server Side Docker are available here.
As also shown in Figure 2 you can also check the option “Run GPRO server locally using Docker” to let you to automatically start the GPRO container each time you run RNASeq (Also note that if you have this option checked you do not need to type the port). You can test if the app is connected to the Server Side clicking on the tab “Test connection settings”. Alternatively, if you install the Server Side manually (without the Docker) just add the IP of the remote server where the Server Side is hosted, add the port information (by default 22) and keep the Option “Run GPRO server locally using Docker” unchecked.
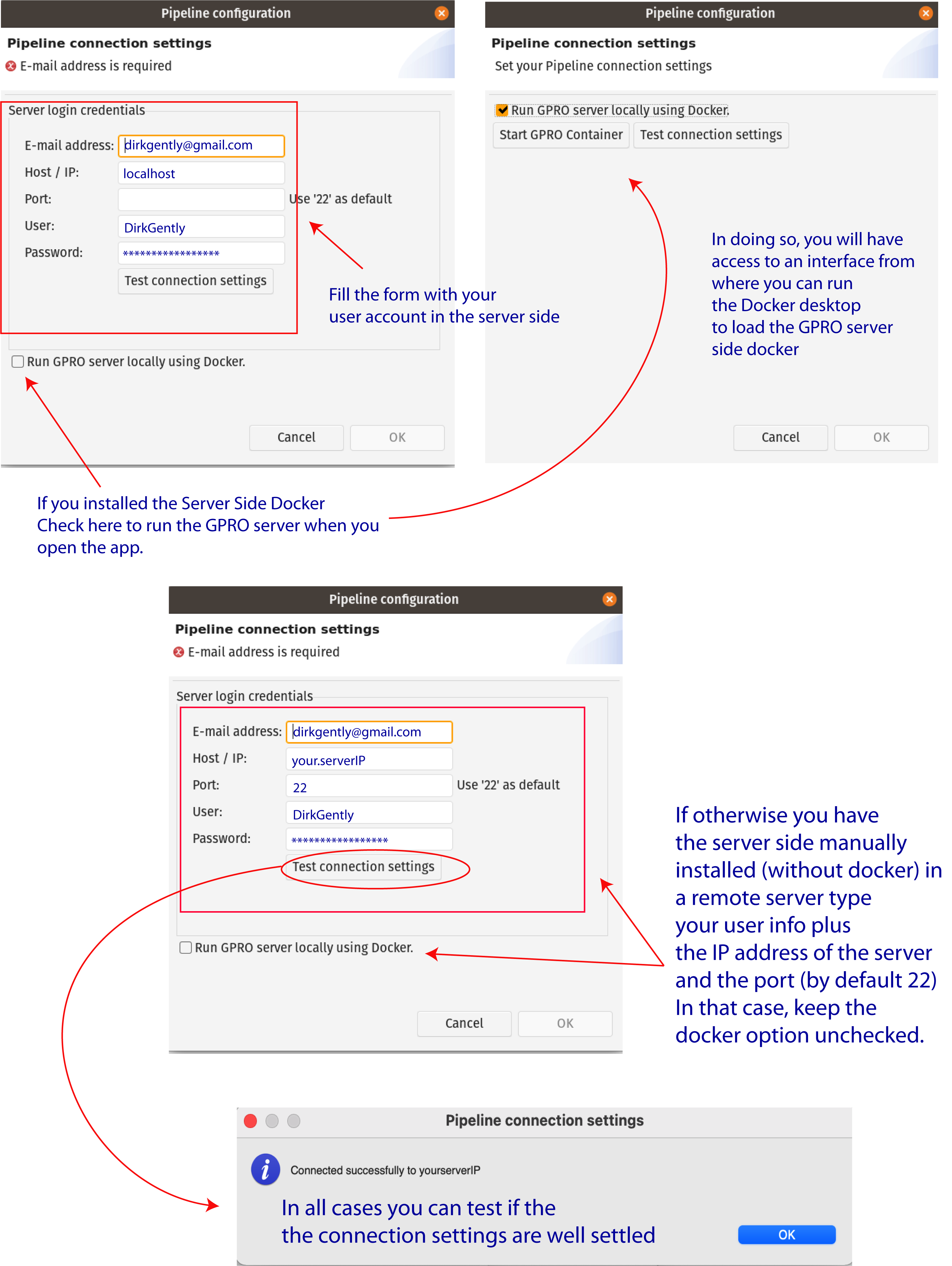
Figure 2: Server connection dialog.
[RNASeq.app → Show package contents → Contents → MacOS → RNASeq.ini.].
Xms1024m (Minimum allocated memory)
|
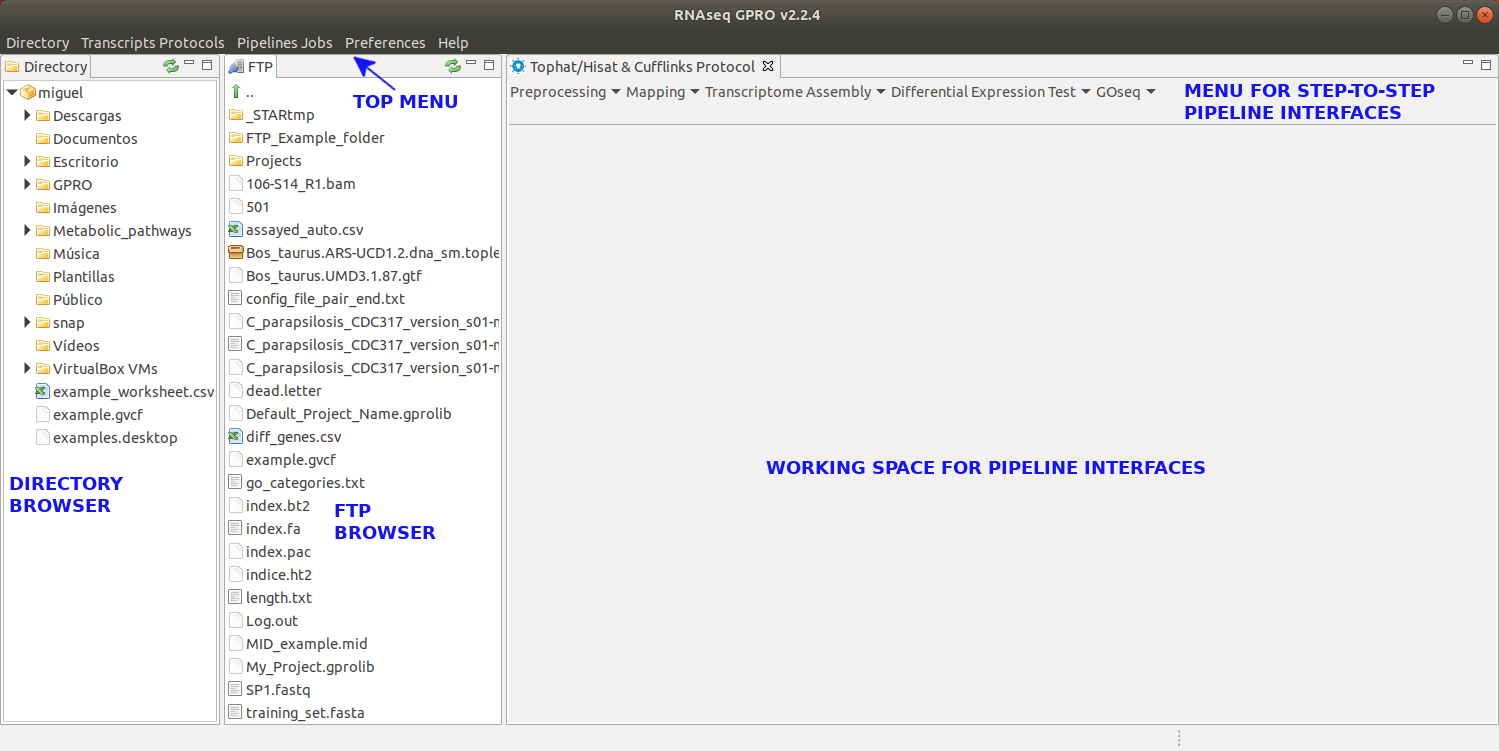
Figure 3: Main layout of RNASeq. Both the Directory and FTP Browser windows can be either resized or masked by clicking on the window icons at the top right corner of their respective windows. All files and folders contained in either of these browsers can be managed manually using the mouse. Please keep in mind that the window views shown in this manual will change depending on the operative system used.
[Directory → Select directory folder ] : Selects a workspace from the directory browser.[Directory → Show ] : Shows the Directory Browser.[Directory → Hide ] : Hides the Directory Browser.[RNAseq Protocols → Pipeline Mode] : Configuration view to set up your RNASeq experiment and launch a RNASeq pipeline.[RNAseq Protocols → Step-by-Step Mode → Tophat & Cufflinks Protocol] : View of the Tophat & Cufflinks Protocol in manual mode.[RNAseq Protocols → Step-by-Step Mode → Mapping & Counting Protocol] :View of the Mapping & Counting protocol in manual mode.[Pipeline Jobs → Jobs Tracking System ] : For jobs tracking.[Pipeline Jobs → FTP Transfers ] : screen for tracking the jobs of the sFTP protocol. [Preferences → Pipeline Connections Settings] : Gives access to the server login details setup your user credentials for accessing the server.[Help → About RNASeq ] : Technical details and copyright of RNASeq, as well as links to the license and user manual.
Figure 4: Example of interface for a CLI tool provided by the ”RNASeq” Step-By-Step mode. The figure shows the interface for Cufflinks.
The procedure is illustrated in Fig.4. Basically, you only need to select with the mouse the input file/s in FTP browser and drag it/them to their respective fields of the interface. The same can be done for the folder/s where you would like to have declared the output. Next, and if any, fill all other mandatory field(s) in the input/output section. If an input field is invalid or missing, you will get an error icon ![]() beside the field (to see the error message you will need to hover the mouse over that icon). Next, configure the form for options and parameters available in the second section. If you set a parameter out of the possible range, you will receive a warning icon
beside the field (to see the error message you will need to hover the mouse over that icon). Next, configure the form for options and parameters available in the second section. If you set a parameter out of the possible range, you will receive a warning icon ![]() . IIf you need more information about any input or parameter, click on the exclamation mark ! in front of the input field name.
. IIf you need more information about any input or parameter, click on the exclamation mark ! in front of the input field name.
Filling the section of options and parameters is not mandatory. If you run the program without inputting any parameters and/or condition, RNASeq will execute this program using the default conditions for this program.
Once the job input fields and the parameters have been fulfilled and/or configured, you only need to click to start button at the end the interface form to run the Job. If the job has been successfully launched, you will get a confirmation message. Otherwise revise again all input and outputs upload and the options if any.





